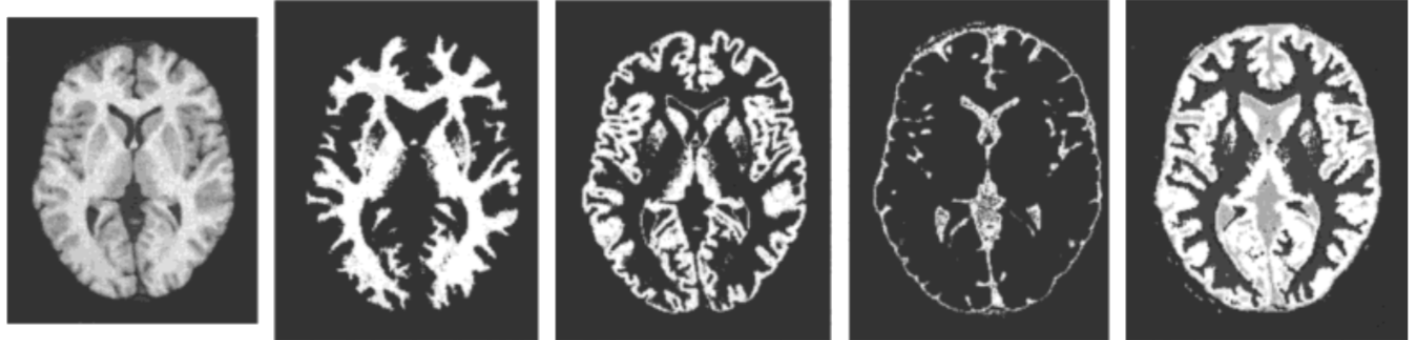
Topology correction of brain surfaces
Brain surfaces provide a reliable representation for cortical mapping. The construction of correct surfaces from magnetic resonance images (MRI) segmentation is a challenging task, especially when genus zero surfaces are required for further processing such as parameterization, partial inflation and registration. The generation of such surfaces has been approached either by correcting a binary image as part of the segmentation pipeline or by modifying the mesh representing the surface. During this task, the preservation of the structure may be compromised because of the convoluted nature of the brain and noisy/imperfect segmentations. In this paper, we propose a combined, voxel and surface based, topology correction method which preserves the structure of the brain while yielding genus zero surfaces. The topology of the binary segmentation is first corrected using a set of topology preserving operators applied sequentially. This results in a white matter/gray matter binary set with correct sulci delineation, homotopic to a filled sphere. Using the corrected segmentation, a marching cubes mesh is then generated and the tunnels and handles resulting from the meshing are finally removed with an algorithm based on the detection of nonseparating loops.
The approach was validated using 20 young individuals MRI from the OASIS database, acquired at two different timepoints. Reproducibility and robustness were evaluated using global and local criteria such as surface area, curvature and point to point distance. Results demonstrated the method capability to produce genus zero meshes while preserving geometry, two fundamental properties for reliable and accurate cortical mapping and further clinical studies.
Topographic Analysis of Electrode Contacts in Human Cortical Stimulation
Electric chronic stimulation of the human motor cortex (ECSM) has been reported to alleviate chronic severe pain. However the mechanism of action of ECSM is still hypothetical. This is due mainly to the poor knowledge of, 1) the electric diffusion through the multiple structures beneath the epidural contacts (i.e. dura matter, cerebrospinal fluid space, arachnoid membrane, grey and white matter layers, pie mere and vascular tree), 2) the absence of consensus concerning the stimulation parameters (mono versus bipolar stimulation, cathodic or anodic current) and 3) the detailed cortical topography of the contacts. In this study we focused on the precise identification of the cortical areas covered by the electric contacts in a series of twelve patients operated on for ECSM. We proposed a new automatic tool for topographic analysis able to compute 2D maps from the 3D anatomic MRI with bijective transformation (point-to- point correspondance). Anatomical regions of interest (AROIs) were visually identified, manually outlined and extracted (Iplan, BrainLab, Germany) for further analysis : 1) for the anatomic structures, on pre operative T1-weigthed magnetic resonance imaging (MRI), the frontal (superior or F1, intermediate or F2 and inferior or F3), the pre central and the post central gyrus ; 2) for the electrode contacts (Resume, Medtronic, USA), on post operative computerized tomography (CT). After getting white and gray matter membership maps by automatic segmentation, we produced a cortical mask to build a triangular mesh. We defined a homeomorphism between the 3D mesh and a subset of R2 and could apply in consequence the circle packing algorithm. We built depth maps (distance to the skull), distance to- contact maps (distance to a given electrode contact) and anatomic structure maps.
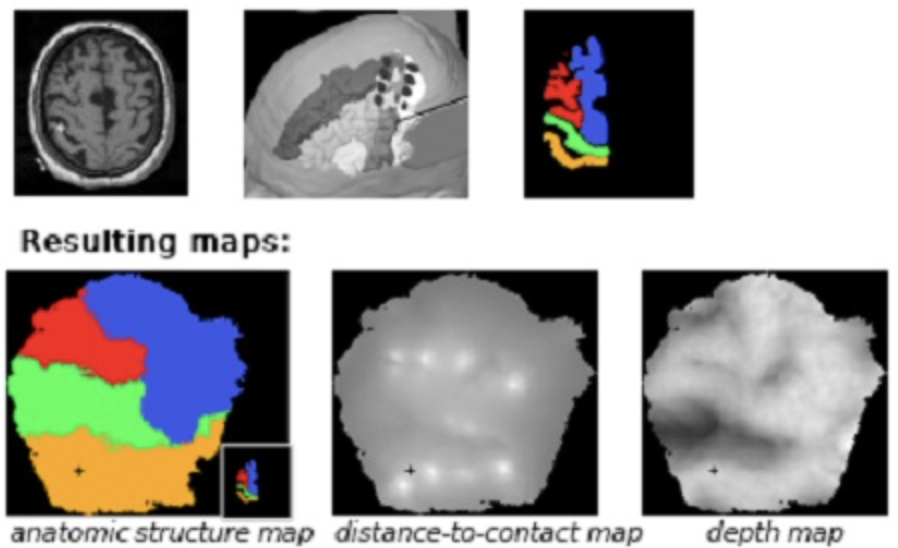

Results showed that it was easier to accurately define the location of the contact projection on the cortex allowing physicians to correlate the benefit with the topography. In particular, because of the unfolding, it was easier to integrate the cytoarchitectonics (i.e. the manually identified AROIs) knowledge in the analysis. Beyond the better understanding of ECSM and indirectly of the pathophysiologic process of chronic pain, this new tool might be used in the future for image guided electrode positioning.
Transcranial Magnetic Stimulation
Transcranial Magnetic Stimulation (TMS) is a technique for brain stimulation. As another techniques like ElectroConvulsive Therapy (ECT) or the implantation of electrodes into motor cortex, TMS excites neurones. But contrary to those techniques, TMS is painless and noninvasive. Indeed, neurones are excited by electrical current induced by a rapidly changing magnetic field. This field is created by the discharge of thyristor (during about 300 μs) into a coil i.e a copper winding surrounded by a water-cooled system. The current circulating through the coil is about 10000 A in order to allow magnetic field to cross the skull of the patient.
Using TMS is sometimes still empirical. The clinician who wants to stimulate a specific cortex area given an expected result (phosphene, modification of vision, motor effect) will perform several stimulations, scanning over the patient’s scalp to find the most important response.
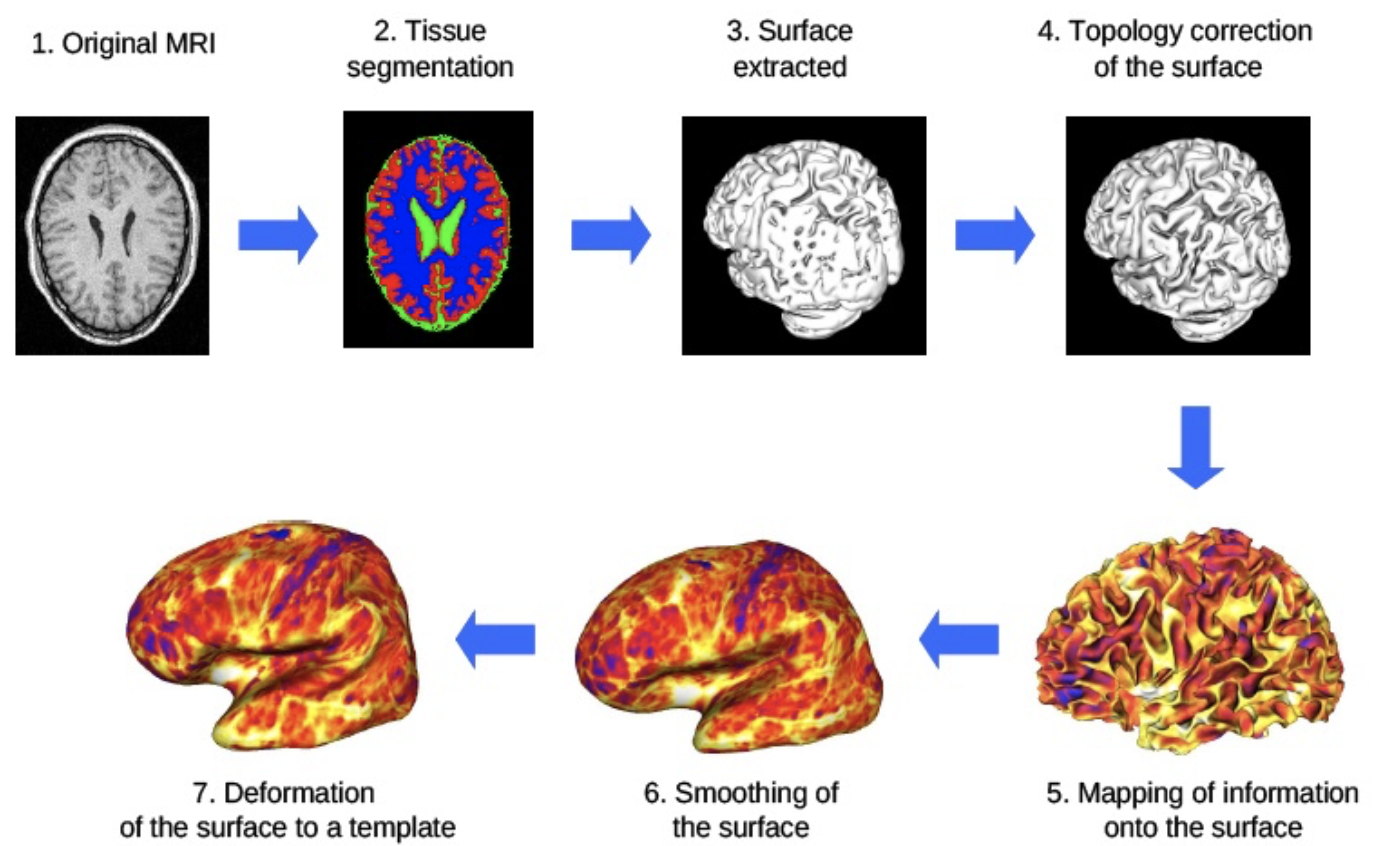
We thus proposed in this project to develop a transcranial magnetic stimulation simulator : according to position and orientation of the coil and to the parameters of the stimulation, the simulator will give a modelling of the stimulation effects. Result will be mapped on MRI image of the patient.
The TMS simulator is composed of three main tools :
- A numerical tool, to compute magnetic field generated by a given coil at any point of the empty-space. The aim is first to assess the magnetic field induced by the coil, then to provide a finite-element scheme boundaries conditions and finally to finite-element computation in free space.
- A design tool, able to take into account the specificity of each winding copper shape. For a given contour, the user has only to define a 3D-parametrical curve and the expression of its derivative
- An interface tool, providing some results such as the maps of magnetic and potential vector field, mapped on a 3D brain of the patient, and some interactions ( position/orientation of the coil w.r.t. the patient…)
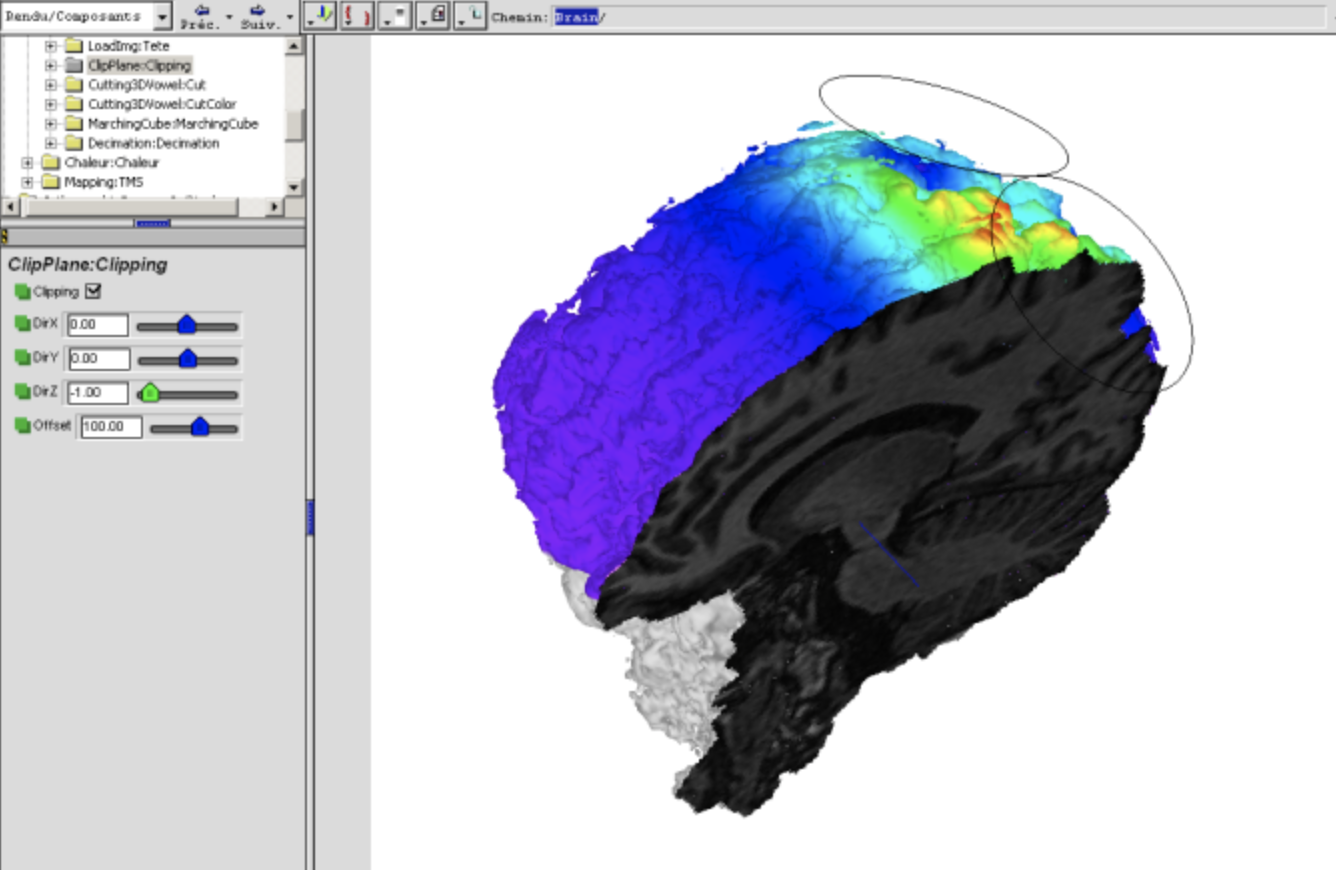
Analysis of Segmentation Methods
The objective of this project was to test new methods and compare state-of-the-art software for brain tissue segmentation and brain morphometry from 3D MR images. The methods we included in our study concerned local algorithms (LIMOS and Bergen), and segmentation algorithms provided by SPM5, FreeSurfer, and FSL. This effort was related to a multidisciplinary project on cognitive aging where the group in Bergen has acquired dual volume T1-weighted 3D brain images from 110 elderly subjects (age : 46-79 years) where also genotyping (APOE, Chrna4, BDNF) and results from extensive neuropsychological phenotyping is obtained. The best performing methods from the brain segmentation and morphometric studies were then be applied to this large image collection and numerical measures incorporated into statistical analysis for better insight in the aging process.
More specifically, the present project addressed the following issues :
- Methods for performance evaluation
- Optimal use of dual MR acquisitions (separate volume classification ? image registration ? value of single versus dual acquisition ?)
- Influence of spatial inhomogeneities and noise in the MR signal, how to correct it ? Derive parameters (regional tissue volumes, cortical thickness, etc) that can be used in a broader statistical analysis with other biological and behavioral data
- Make results (test-data, programs, evaluations) available on Web
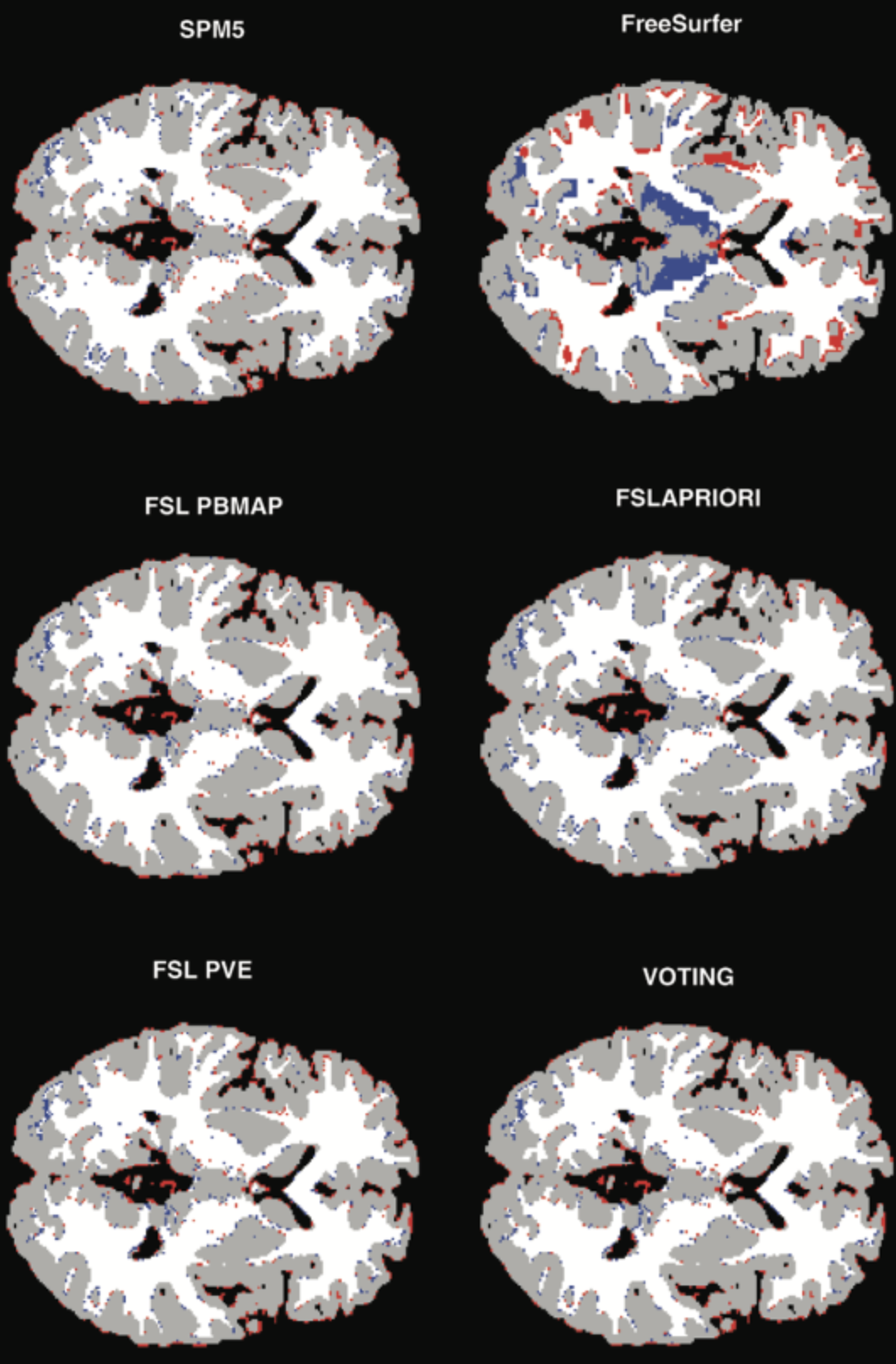
Fusion of Image/Expert information
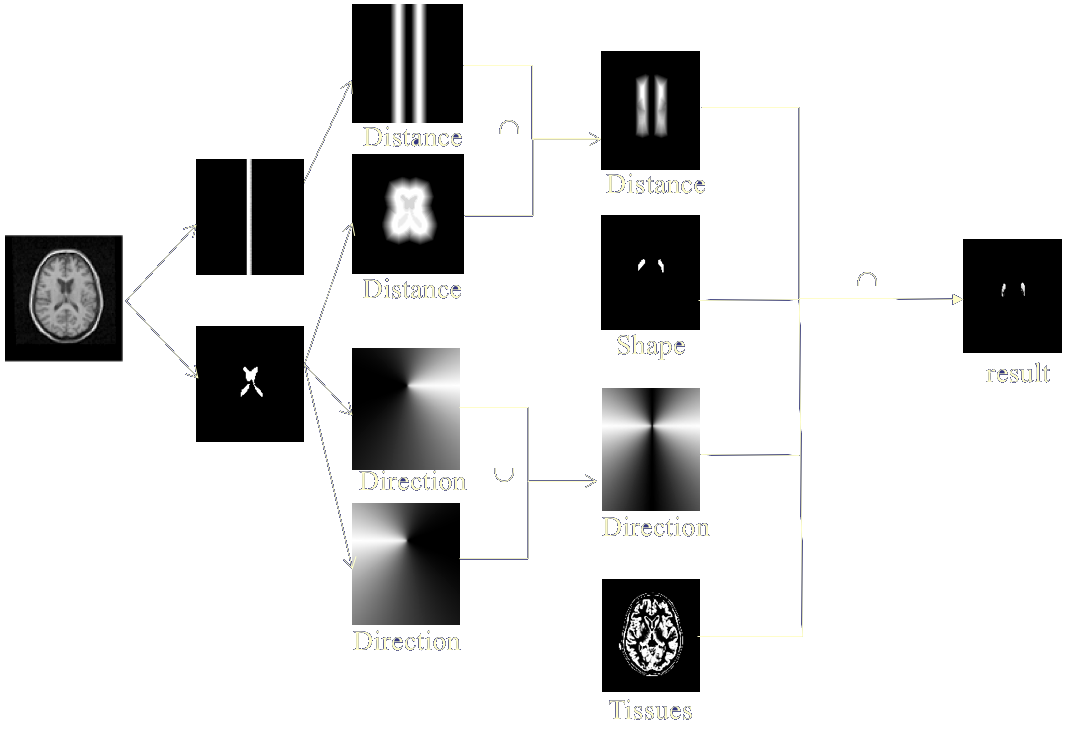
Expert knowledge information relate to indications of position, of form…, and can be clarified as fuzzy proposals (modeling the reflexions of a specialist) and/or as a brain atlas . Fusion thus takes place on heterogeneous and rather complementary data which are not necessarily expressed within the same formal framework (numerical for the images and the atlas, linguistics for expert knowledge). The interest of fusion is here to push back the decision (the segmentation of structures for example) until having sufficient information. This remark, which prevails in general for fusion, is remarkably illustrated in this type of problem, where the information of gray level or coarse position is not enough with the segmentation, but where the aggregation of the whole set of knowledge allows to infer a reliable decision.
The use of this type of method relates mainly to the segmentation of cerebral structures starting from images, atlas and symbolic data modeling expert knowledge. This segmentation can take place for the whole brain with, for example, a research of the hemispheres and cerebellum or stick more particularly to the detection of fine anatomical structures implied in specific pathologies. The clinical implications go then from the targeting of structures for a help to the surgical act to the follow-up of morphometric measurements of a pathological entity.
We applied this method :
- to the segmentation of the subthalamic nucleus within the framework of the treatment of the idiopathic Parkinson’s disease by electrostimulation
- to the segmentation of striatum
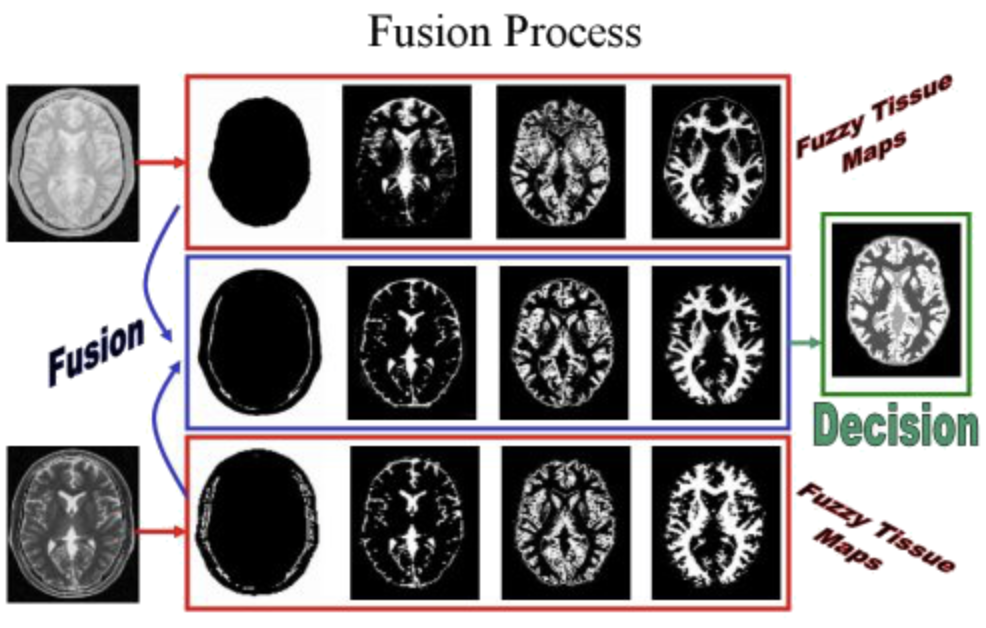
MR/MR Image Fusion
We proposed in this project to aggregate MR images resulting different types of acquisition. The goal here was to precisely quantify volumes of principal cerebral tissues (white substance, gray substance, cerebrospinal fluid). Generally, a precise quantification of these volumes must make it possible to help with the diagnosis, the comprehension and the treatment of some pathologies. We illustrated the concept on the fusion of T2-weighted (T2) and proton density (PD) images. The method was a three step fusion process. The first one involved the computation of fuzzy tissue maps in both images by means of a possibilistic clustering algorithm. The choice of fuzzy logic is motivated by the nature of information. The second step was an aggregation step, while the final segmentation was is the last phase of the process.
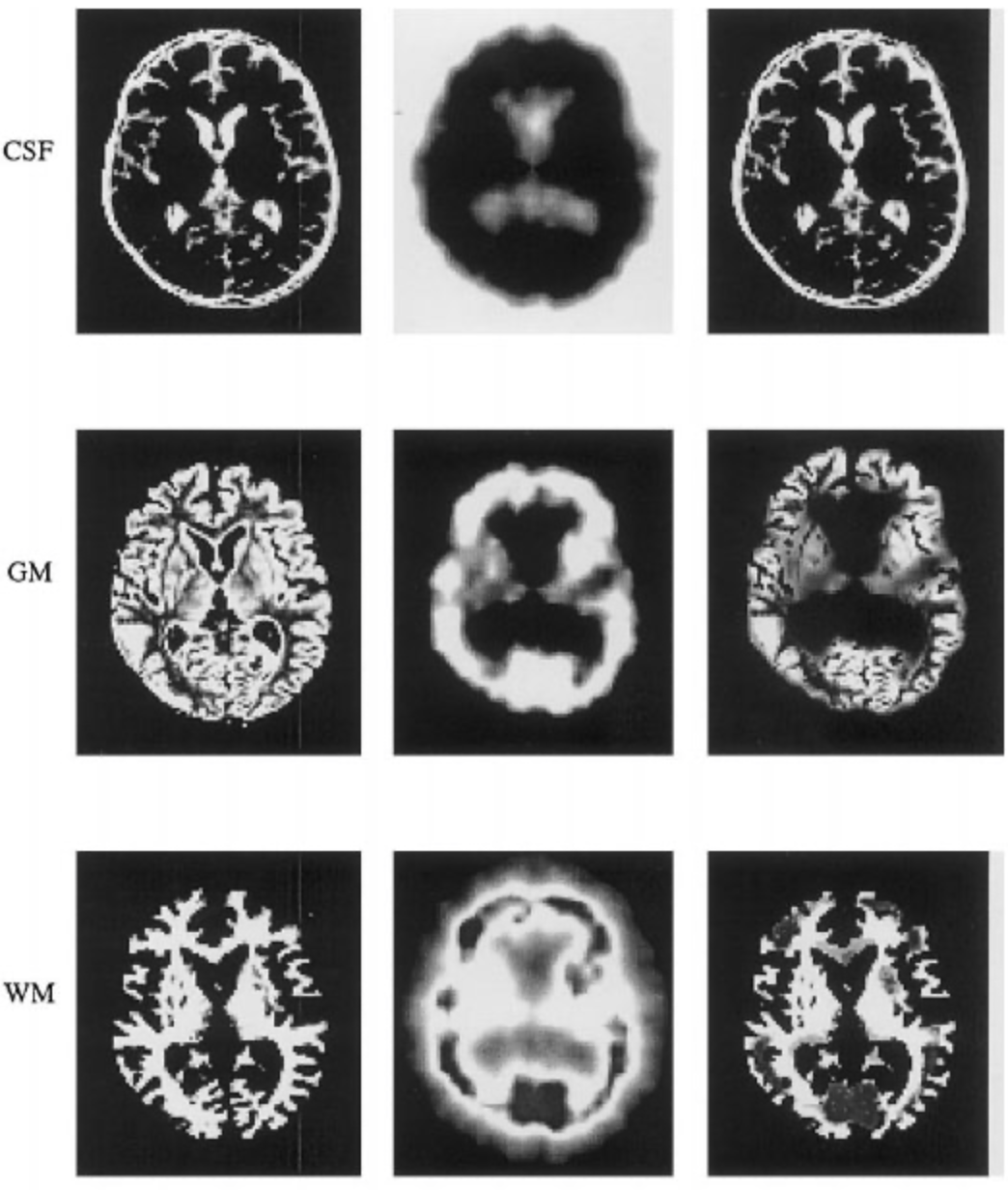
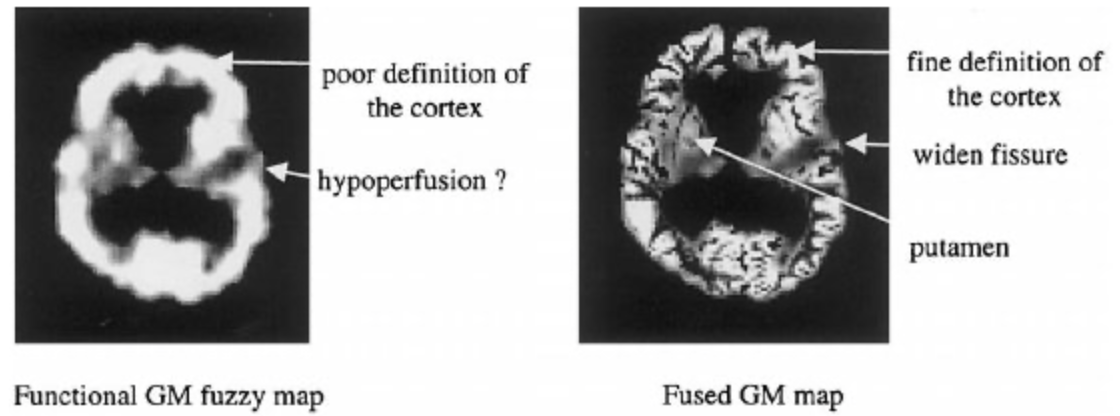
Anatomical / Functional Image Fusion
We proposed to fuse anatomical (A) and functional (F) information coming from medical imaging. We focused on brain imaging, although the principle may be extended to other organs provided image are spatially registered. The anatomical images mainly consisted of MR images that give information on structural composition of the organ (distribution of tissues, fine spatial resolution). They can be obtain using a wide range of parameters, so that the resulting images provide sufficient contrasts between the different structures to be located (tissue interfaces, tumors,…). The functional images were obtained by injecting a radioactive tracer that preferentially characterized a physiologic (e.g., glucose metabolism) or a biologic process (e.g., blood flow). Two major functional modalities are used in brain imaging : SPECT and PET. These images had a bad spatial resolution and are not informative on what they were not supposed to represent.
In the following, anatomical and functional images were supposed to be registered. The same voxel (x,y,z) in the images thus corresponds to the same anatomical location. The registration procedure must be accurate enough to limit the amount of imprecision introduced. We then proposed a three-step fusion process
- Image modeling : information was extracted from the images and was modeled in a common theoretical frame to manage vague and ambiguous knowledge
- Fusion of image information : the information was then aggregated with a fusion operator. This operator must affirm redundancy and manage the complementarities and conflicts that often underline the presence of a pathology ;
- Decision : a fusion image was proposed that synthesized the available information.
Brain image segmentation
The ability of precisely quantifying cerebral tissue volumes in various diseases (Alzheimer, hydrocephaly…) must make it possible to largely improve the diagnosis, the treatment and the general understanding of pathologies. We proposed in 2000 a unsupervised fuzzy classification scheme for brain tissue segmentation. The boundary between brain in an MR image is smooth or fuzzy rather than sharp or crisp. Moreover, noise, partial volume effect and anatomical variations within pure tissue (White Matter,Gray Matter, Cerebrospinal Fluid) activities introduce uncertainty and fuzzyness in the image. We thus used a fuzzy clustering algorithm rather than a crisp one, and particularly the possibilistic clustering algorithm (PCA). PCA allows to interpret memberships as absolute degrees of belonging, whereas they are similar to degrees of sharing or relative memberships in the case of FCM. Because of the complex nature of (GM/WM) or (GM/CF) interfaces in MR images, a significant number of voxels contains a mixture of two or three tissues.